Scientists from the Universities of Manchester, Aberystwyth and Cambridge have created an artificial intelligence, a robot scientist they call Eve, that can discover new drugs rapidly and cheaply.
The team believes Eve will revolutionize new drug basic research and development. They demonstrated the robot’s potential when she discovered, all by herself, that an anti-cancer compound might also be useful in the fight against malaria.
The scientists wrote in the journal Interface that using robot scientists is a natural extension of the increasing automation that is occurring in science.
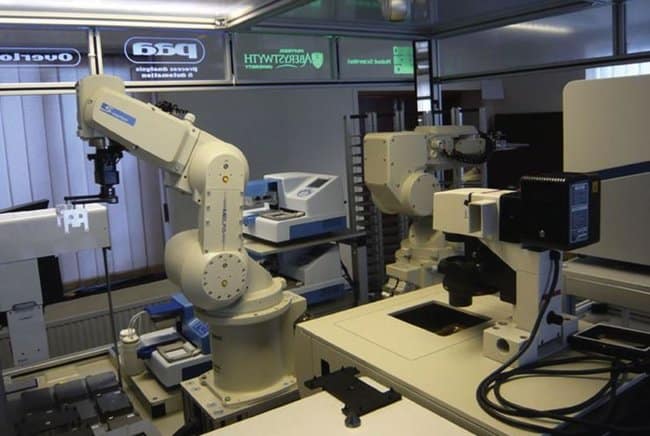

‘Eve’ the robot-scientst can screen more than 10,000 compounds per day. (Photo: Interface)
Eve can automatically develop a hypothesis, test it to explain observations, carry out tests using lab robotics, interpret her findings to adjust her hypothesis, and re-do the cycle “automating high-throughput hypothesis-led research.”
The robot scientist is also competent in recording scientific knowledge. With experiments being conceived and carried out by computer automatically, the artificial intelligence is able to completely capture and digitally monitor every aspect of the scientific process.
Adam came first, and then Eve
Adam, who was created by scientists at the Universities of Aberystwyth and Cambridge, was the first robot scientist to discover new scientific knowledge on its own.
The same team that created Adam have now made Eve. Her purpose is to speed up the drug-discovery process, as well as doing this much more cheaply and within budgets.
According to the scientists, Eve can help spot potential drug candidates for African sleeping sickness, Chagas’ disease, and other neglected tropical diseases, as well as more common ones like malaria.
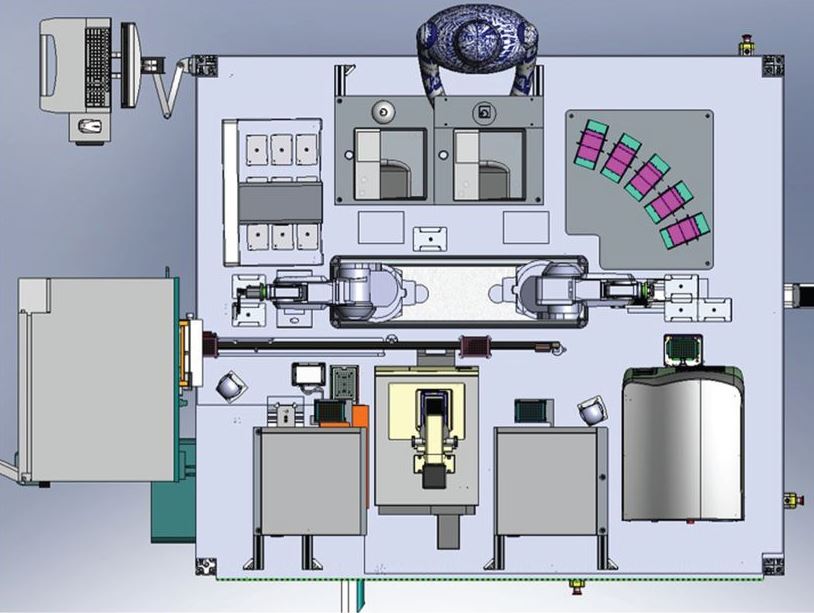
A diagram of Eve, showing the location of the main instruments. (Photo: Interface)
Professor Ross King, from the Manchester Institute of Biotechnology at the University of Manchester, explained:
“Neglected tropical diseases are a scourge of humanity, infecting hundreds of millions of people, and killing millions of people every year. We know what causes these diseases and that we can, in theory, attack the parasites that cause them using small molecule drugs. But the cost and speed of drug discovery and the economic return make them unattractive to the pharmaceutical industry.”
“Eve exploits its artificial intelligence to learn from early successes in her screens and select compounds that have a high probability of being active against the chosen drug target. A smart screening system, based on genetically engineered yeast, is used.”
“This allows Eve to exclude compounds that are toxic to cells and select those that block the action of the parasite protein while leaving any equivalent human protein unscathed. This reduces the costs, uncertainty, and time involved in drug screening, and has the potential to improve the lives of millions of people worldwide.”
With the robot scientist, early-stage drug design is done automatically. She begins by systematically testing every member of a large group of compounds in the standard brute-force way of conventional mass screening.
She screens the molecules against assays designed to be automatically performed. These can be done much faster and for much less money compared to the custom-made assays that are standard today.
Consequently, she can apply more types of assays, use the screening facilities much more efficiently, thus enhancing the profitability of a drug discovery within a designated budget.
Eve can screen more than 10,000 molecules each day
Eve’s robotic system is able to screen in excess of 10,000 molecules daily. However, mass screening is still comparatively slow and burdensome because every compound in the library is tested. The device does not have the intelligence to make use of what it has learnt during the screening process.
In order to make this process faster, Eve chooses a subset of the library randomly to find compounds that pass the first assay. Any hopeful “hits” are re-tested many times to reduce the risk of false positives.
She then takes this set of confirmed “hits” and with the use of machine learning and statistics learns to predict new structures that could score better against the assays. While not currently able to synthesize those compounds, the team says this feature could be added in future versions of Eve.
Steve Oliver, from the Department of Biochemistry, University of Cambridge, said:
“Every industry now benefits from automation and science is no exception. Bringing in machine learning to make this process intelligent – rather than just a ‘brute force’ approach – could greatly speed up scientific progress and potentially reap huge rewards.”
To see whether Eve’s approach might be viable, the researchers developed assays targeting key compounds from parasites that cause malaria, Chagas’ disease and schistosomiasis, and tested a library of approximately 1,5000 clinically approved compounds against them.
On her own, Eve discovered that a molecule that had been previously researched as an anti-cancer drug inhibited DHFT, a key molecule in the malaria parasite. She found a likely new therapeutic application for the existing compound.
DHRF-inhibiting drugs are given to millions of children to protect them from malaria. Unfortunately, new strains of the parasite have evolved which are resistant to current medications, thus making the discovery of new drugs increasingly more urgent.
Professor Oliver said:
“Despite extensive efforts, no one has been able to find a new antimalarial that targets DHFR and is able to pass clinical trials. Eve’s discovery could be even more significant than just demonstrating a new approach to drug discovery.”
The project was funded by the Biotechnology & Biological Sciences Research Council and the European Commission.
Reference: Kevin Williams, Elizabeth Bilsland, Andrew Sparkes, Wayne Aubrey, Michael Young, Larisa N. Soldatova, Kurt De Grave, Jan Ramon, Michaela de Clare, Worachart Sirawaraporn, Stephen G. Oliver, Ross D. King. Interface. Published 4 February 2015. DOI: 10.1098/rsif.2014.1289. “Cheaper faster drug development validated by the repositioning of drugs against neglected tropical diseases.”
Video – Artificial Intelligence
When computer scientists use the term AI (artificial intelligence), they are referring to software that makes smartphones, computers, robots, and other devices smart. In other words, their thought processes and behaviors are similar to ours (humans’).

